Cloud proteomics platforms are increasingly shaping how laboratories manage, analyze, and share large-scale proteomics datasets. Advances in high-resolution mass spectrometry have dramatically increased the volume and complexity of proteomics data, creating new challenges for storage, computational analysis, and collaboration.
Cloud proteomics platforms offer a framework for scalable data processing and distributed collaboration by integrating cloud computing infrastructure, shared repositories, and computational workflows. These systems enable laboratories to handle large datasets generated from proteomic studies while supporting reproducibility, accessibility, and cost management across research teams.
Cloud computing infrastructure for proteomics data management
Modern proteomics experiments—particularly those using high-throughput mass spectrometry—generate terabytes of raw spectral data. Managing this data locally can place substantial strain on laboratory storage infrastructure and computational resources.
Cloud computing environments provide elastic storage and computing capacity (Table 1), allowing laboratories to scale resources according to experimental demand.
Table 1: How cloud proteomics platforms compare to local infrastructure.
Cloud Proteomics Platforms
Limited by local hardware
Scalable, on-demand storage
Computational resources
Elastic compute clusters
Manual transfer between labs
Centralized collaborative access
Requires dedicated IT management
Hardware upgrades required
Cloud-based environments also support integration with existing laboratory pipelines. Raw data produced by mass spectrometry instruments can be uploaded to centralized cloud storage where it can be processed by computational pipelines without requiring local workstation resources.
These capabilities allow laboratories to expand analytical capacity without maintaining large local compute clusters, which can reduce operational complexity.
Proteomics data sharing and the role of cloud-based repositories
Proteomics research increasingly relies on open data sharing to support transparency, reproducibility and cross-study analysis. Cloud-based data repositories provide standardized platforms where datasets can be deposited, accessed, and reanalyzed by the scientific community.
Examples include initiatives such as the ProteomeXchange Consortium and the widely used PRIDE Archive, which coordinate the submission and dissemination of proteomics datasets across multiple repositories.
Cloud proteomics platforms facilitate data sharing by enabling:
Many repositories also adopt FAIR data principles—ensuring that datasets are findable, accessible, interoperable, and reusable.
Standardized metadata improves interoperability between laboratories and enables large-scale meta-analysis across datasets, which is increasingly important in systems biology and biomarker discovery.
Key metadata components often include:
Computational workflows and mass spectrometry integration
The analytical workflow in proteomics is tightly linked to mass spectrometry instrumentation. Cloud proteomics platforms support these workflows by integrating computational pipelines that can process raw spectral data into biological insights.
Typical mass spectrometry data analysis pipelines involve multiple stages:
-
Raw spectral data acquisition
-
Peak detection and feature extraction
-
Peptide identification via database searching
-
Protein inference and quantification
-
Statistical analysis and biological interpretation
Cloud environments enable these workflows to run across distributed computing resources, which can significantly accelerate processing times for large datasets.
Common workflow advantages include:
Cloud-based analysis platforms are particularly useful in studies involving large cohort proteomics datasets, multiomics integration, high-throughput biomarker discovery, or cross-institutional collaborative research (Figure 1).
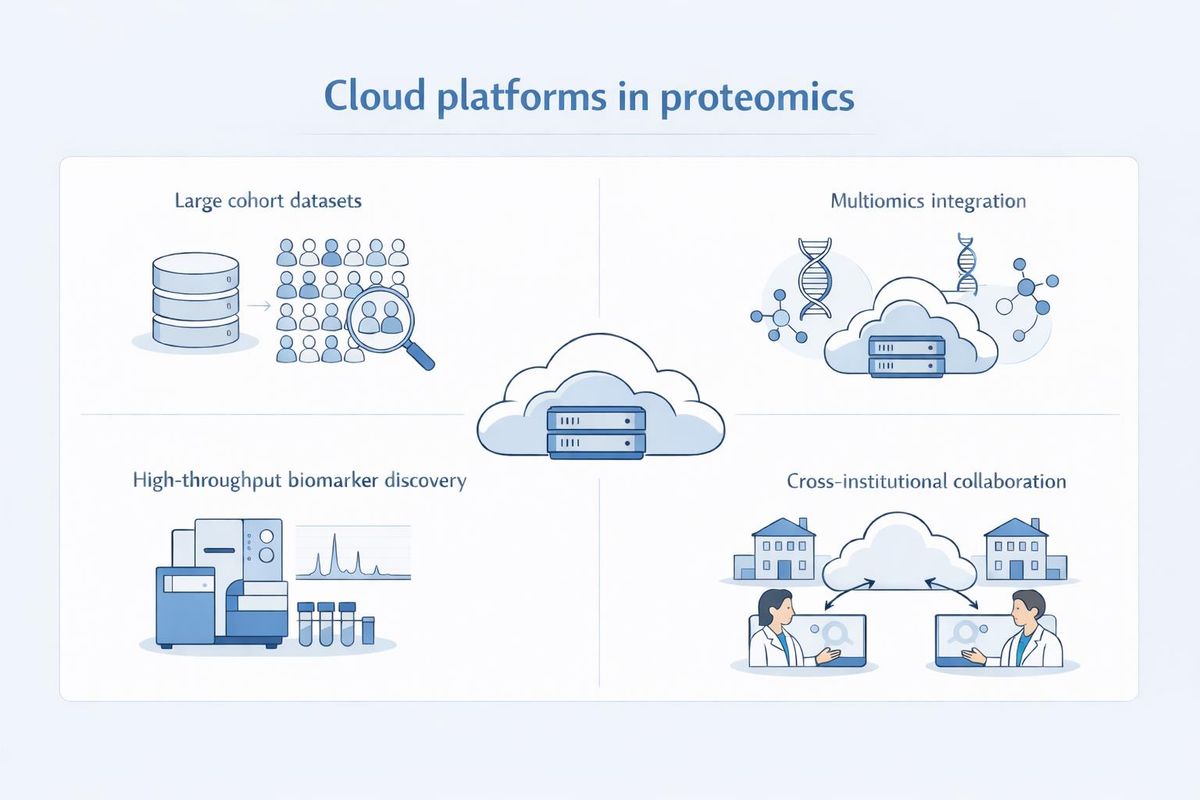
Figure 1: How cloud-based platforms support proteomics research. Credit: AI-generated image created using Microsoft Copilot (2026).
By distributing computational workloads across multiple processors, cloud computing enables efficient processing of large spectral libraries and complex search spaces.
Cost optimization strategies in cloud proteomics platforms
Although cloud infrastructure provides scalability, laboratories must carefully manage resource allocation to control operational costs. Cost optimization strategies focus on balancing computational demand with available infrastructure resources.
Key strategies commonly used in cloud-based proteomics workflows include:
Cost considerations often involve evaluating several factors, including data storage volume, the frequency of data access, the computational intensity of analysis pipelines, and data transfer requirements between institutions.
A common approach is tiered storage architecture, which separates data into categories (Table 2).
Table 2: How storage tiers in cloud-based proteomics platforms are used.
High-performance storage
Frequently accessed datasets
Long-term dataset preservation
This model allows laboratories to maintain large proteomics datasets while minimizing ongoing storage costs.
Additional optimization strategies include:
When combined with scalable infrastructure, these strategies can support long-term sustainability of cloud-based proteomics workflows.
Data security, compliance, and governance in cloud proteomics platforms
Data governance is an important consideration when using cloud infrastructure for biological research. Proteomics datasets may include clinical samples or patient-derived data, which require secure handling and regulatory compliance.
Cloud proteomics platforms typically incorporate several security features:
Many research institutions also implement data governance policies that define how datasets are shared, stored, and archived across collaborative projects. Proper governance ensures that cloud-based data sharing can occur while maintaining data integrity and regulatory compliance.
Key governance considerations include:
How cloud proteomics platforms are shaping data sharing in research
Cloud proteomics platforms are transforming how laboratories manage and analyze large-scale proteomics datasets. By combining scalable computing infrastructure, standardized data repositories, and integrated computational workflows, these platforms enable researchers to process complex datasets while supporting collaborative research.
The integration of cloud computing with mass spectrometry workflows allows laboratories to handle increasing data volumes without maintaining extensive local computing infrastructure. At the same time, standardized repositories and FAIR data principles promote transparency and reproducibility across proteomics studies.
As proteomics experiments continue to grow in scale and complexity, cloud-based platforms are likely to play a central role in enabling efficient data sharing, scalable analysis, and collaborative discovery in laboratory research environments.