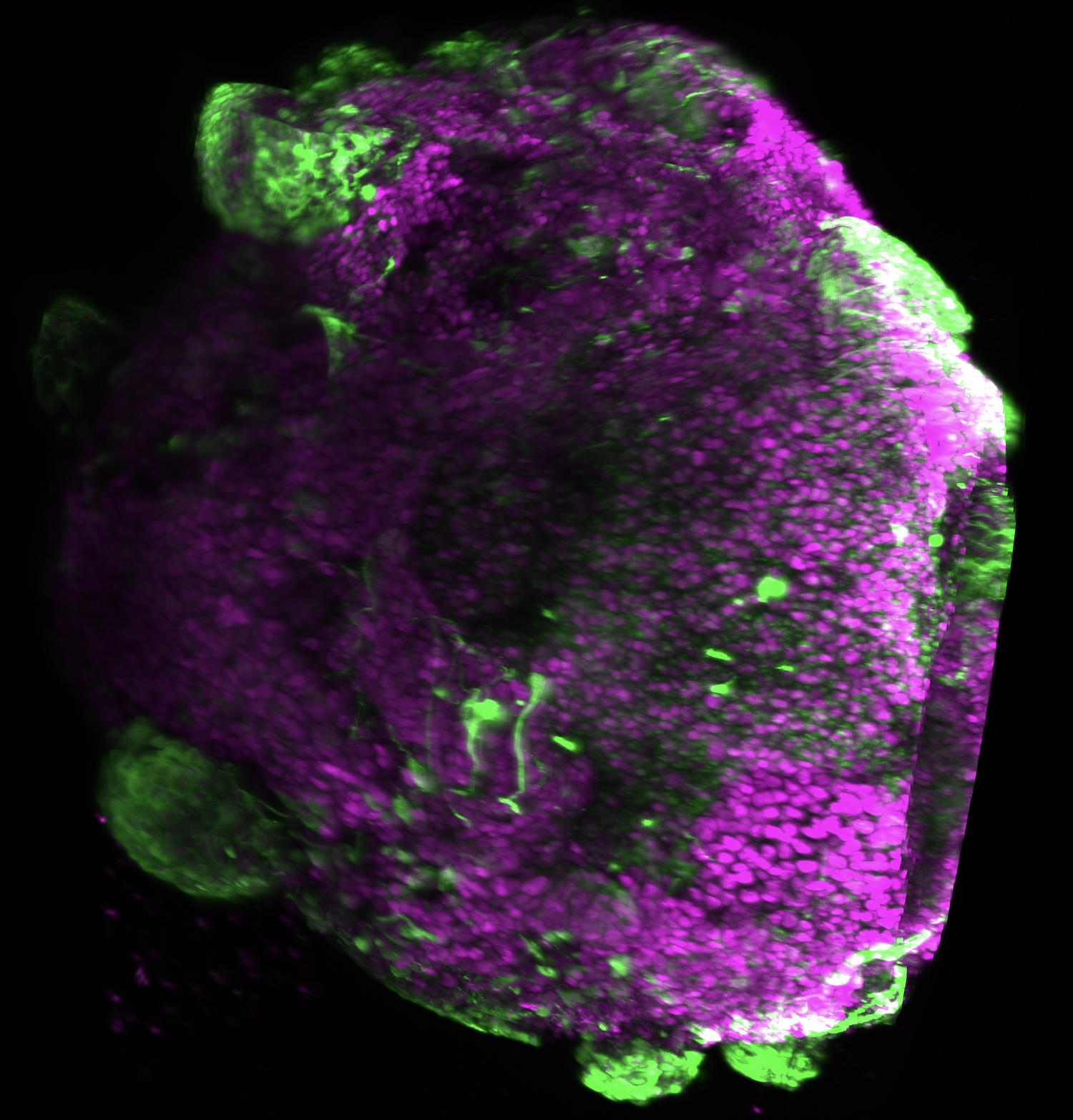
Cortical organoids have been the closest thing to human cortex in a dish for years, but they don't develop regional identity. No recognizable motor cortex, visual cortex, or prefrontal cortex inside an organoid. So whichever region-specific brain disease you'd want to model (motor-cortex ALS, prefrontal FTD, sensory-cortex disease), you can't really hit a region. That bottleneck has limited a decade of organoid research.
A lab at the University of Alabama Birmingham is testing a fix that adapts 1990s developmental biology. To map how the embryo gets its body plan, biologists used to implant beads soaked in signaling molecules into chick embryos and watch cellular identity shift with bead position. The new experiment applies the same trick to human cortical organoids: opposing FGF-2 and Activin-A beads on agarose pedestals around the organoid, recreating the morphogen gradients that pattern cortex during development.
If this protocol works, three things open up:
Region-specific disease modeling at scale. The KOLF2.1J iPSC line used here is the same one the NIH iNDI initiative built CRISPR-edited disease panels on top of (ALS, Parkinson's, AD, FTD, HD lines), so disease-allele organoids are the natural follow-up.
Biological-substrate computing. Platforms like Cortical Labs DishBrain currently train on cortical-organoid tissue that isn't actually structured like cortex. A patterned organoid with real regional identity changes the kind of computation you could train.
Methods generalization. The bead-and-pedestal approach uses off-the-shelf reagents and a standard plate format, so unlike microfluidic gradient generators, it could actually leave the originating lab.
Curious which of these futures matters most to people here, and whether you'd bet on the methods generalizing.
https://www.researchhub.com/proposal/7659/engineering-morphogen-gradients-to-grow-larger-spatially-patterned-human-cortical-organoids
3 Comments
The endeavor to instill regional identity within human cortical brain organoids by introducing opposing growth factor beads represents a sophisticated attempt to bridge the gap between static biological models and the dynamic complexity of living tissue. Historically, the challenge with lab grown neural structures has been their tendency to develop as a relatively uniform mass of cells, lacking the precise spatial organization and distinct functional zones found in a natural brain. By revisiting embryological principles established decades ago and applying them to modern stem cell cultures, researchers are attempting to recreate the fundamental chemical gradients that guide a developing embryo. This process functions as a mechanical blueprint, where the placement of specific biochemical signals acts as a grounding rod to direct the flow of cellular development into organized, recognizable regions such as the forebrain or midbrain.
This experimental framework relies on the concept of positional information, where the concentration of a signaling molecule tells a cell exactly where it is in space and what it should become. When beads soaked in opposing growth factors are placed at either end of a three dimensional organoid, they create a field of overlapping chemical signatures that force a systemic transition within the culture. The cells no longer exist in a state of generic potential but are instead compelled to specialize based on their proximity to these specific anchors. This creates a phase shift within the biological system, moving it from a state of disorganized growth toward a purely positive version of anatomical accuracy. It is a deliberate orchestration of nature’s own developmental language, using physical tools to nudge the biological hardware into a more sophisticated state of existence.
The successful implementation of these regional identities would mark a profound evolution in how we study human neurology and disease. By creating organoids that possess a literal and accurate geography, scientists can observe the interactions between different brain regions with a level of clarity that was previously impossible. This grounded approach to modeling life acknowledges that the brain’s power comes not just from the presence of neurons, but from the specific, localized ways those neurons are organized and connected. As these lab grown systems begin to mirror the structural integrity of a real human cortex, the boundary between artificial model and biological reality begins to dissolve, offering a more honest and integrated medium for understanding the complexities of human consciousness and the architecture of the mind.
Submission statement: This experiment sits at the intersection of three future-relevant threads, and I’d like to hear which one matters most to people here.
Most immediate future: region-specific brain-disease modeling at human-cell scale. Motor-cortex ALS, prefrontal FTD, and sensory-cortex disease have all been waiting for a substrate that actually has the regions. Combine that with the existing iNDI CRISPR-edited disease panels on the same iPSC line, and you get a generalizable disease-modeling platform.
Further out: biological-substrate computing. Platforms like Cortical Labs DishBrain currently train on tissue that has cortical neurons but isn’t structured like cortex. A patterned organoid changes what kind of computation you could train and read from.
Furthest out: the methods themselves. The bead-and-pedestal approach uses off-the-shelf reagents and a standard plate format. If it generalizes, organoid morphogen patterning leaves the originating lab in a way microfluidic gradient generators never did.
About time I have been saying this since Covid started.
It’s so obvious yet I could never get the funding